Abstract
Purpose
We aimed to identify the genetic cause to a clinical syndrome encompassing hypohidrosis, electrolyte imbalance, lacrimal gland dysfunction, ichthyosis, and xerostomia (HELIX syndrome), and to comprehensively delineate the phenotype.
Methods
We performed homozygosity mapping, whole-genome sequencing, gene sequencing, expression studies, functional tests, protein bioinformatics, and histological characterization in two unrelated families with HELIX syndrome.
Results
We identified biallelic missense mutations (c.386C>T, p.S131L and c.2T>C, p.M1T) in CLDN10B in six patients from two unrelated families. CLDN10B encodes Claudin-10b, an integral tight junction (TJ) membrane-spanning protein expressed in the kidney, skin, and salivary glands. All patients had hypohidrosis, renal loss of NaCl with secondary hyperaldosteronism and hypokalemia, as well as hypolacrymia, ichthyosis, xerostomia, and severe enamel wear. Functional testing revealed that patients had a decreased NaCl absorption in the thick ascending limb of the loop of Henle and a severely decreased secretion of saliva. Both mutations resulted in reduced or absent Claudin-10 at the plasma membrane of epithelial cells.
Conclusion
CLDN10 mutations cause a dysfunction in TJs in several tissues and, subsequently, abnormalities in renal ion transport, ectodermal gland homeostasis, and epidermal integrity.
Similar content being viewed by others
Introduction
Located in the most apical position of the lateral epithelial cellular membrane, the tight junctions (TJs) selectively control the passage of solutes and water through the lateral paracellular space (known as barrier function),1 maintain cell polarity, and play a role in signal transduction (known as fence function).2 The claudins are integral membrane proteins involved in the formation of the TJ plates with at least 27 members in the annotated mammalian genomes.3, 4 The prominent role of claudins is to determine paracellular selectivity and permeability to small ions by acting as pores or barriers for ions.5 Mutations in CLDN16 and CLDN19 are responsible for familial hypomagnesemia with hypercalciuria and nephrocalcinosis (MIM 248250 and 248190).6, 7 Mutations in CLDN1 cause neonatal sclerosing cholangitis with ichthyosis (MIM 607626),8 and mutations in CLDN14 are responsible for a form of autosomal recessive nonsyndromic deafness (DFNB29, MIM 614035);9 all are autosomal recessive traits. We report a new syndrome affecting several epithelia related to biallelic CLDN10 mutations.
Materials and methods
Patients
All patients and relatives gave their or their legal guardian’s written informed consent. Baseline values were measured after an overnight fast, in sitting position and after withdrawal of any interfering treatment with the exception of oral potassium supplements.
Ethical considerations
This study was approved by the local institutional review board at Hôpital Necker–Enfants Malades, Paris, France, and by the institutional review board of the Shafallah Medical Genetics Center (later QBRI-MGC), Doha, Qatar, both operating according to the declaration of Helsinki.
Light microscopy
Skin biopsy specimens were fixed in 10% formalin, embedded in paraffin, and processed routinely. Sections 3 μm thick were stained with hematoxylin and eosin and examined by light microscopy at different magnifications.
Gene mapping by homozygosity
The two affected individuals from the first sibship of family A, their three unaffected sisters, and their parents participated in the study. DNA was extracted from whole blood using standard procedures. A genome-wide scan for linkage was performed by genotyping each participating member with the Human OmniExpress chips (about 700,000 single-nucleotide polymorphisms) from Illumina (San Diego, CA). The genotyping data and allele calling were carried out by Genome Studio 2011 v.1. The regions of homozygosity were determined using the public access software HomozygosityMapper.10, 11 The software output is linked to a gene search engine GeneDistiller.12
Mutation detection
We designed primers for the amplification of CLDN10 using the Primer3 software (http://frodo.wi.mit.edu/) for the sequencing of all coding exons and the flanking sequences. DNA sequencing was performed utilizing BigDye terminator chemistry (Life Technologies, Carlsbad, CA, USA).
Protein bioinformatics
There are two Claudin-10 splice variants that differ only in their first exon and give rise to two main Claudin-10 isoforms, Claudin-10a and Claudin-10b.13 We worked primarily with Claudin-10b, the isoform that is expressed ubiquitously, and checked if similar results were obtained for Claudin-10a. To determine the extent of the transmembrane helices, we extended both ends of each predicted transmembrane helix to the first charged residue. The serine residue at position 131 (position 129 in Claudin-10a) is located in the C-terminal half of TM3. For the bioinformatic analysis, we obtained a 3D model of the structure of Claudin-10b based on the 3D crystal structure of mouse claudin-15.14, 15 The bioinformatics methodology is detailed in the Supplementary Appendix online.
Immunohistochemistry
Claudin-10 (1/25, polyclonal; Abcam, Cambridge, UK) and filaggrin (1/500, polyclonal; Biolegend, Dedham, MA) antibodies were analyzed by immunohistochemistry in skin biopsy specimens from the patients and normal-skin controls.
The immunohistochemistry reaction was performed on 3- to 4-μm-thick formalin-fixed paraffin-embedded sections. After the deparaffination steps, reactions were carried out according to the automatized system Bond (A. Menarini Diagnosis System, Florence, Italy). Staining was performed with the Menarini refine detection kit.
Furosemide infusion test
After two 30-minute control periods, a priming dose of furosemide (5 mg) was injected intravenously, and then followed by an infusion of 5 mg/hour in 0.9% saline solution at constant rate over 2 hours. After a single 1-hour equilibration period, two 30-minute urine collections were performed. In all periods, urinary creatinine, Na, Cl, Ca, and Mg, and serum creatinine, Na, Cl, Ca, and Mg, were measured.
Analytical methods
Plasma creatinine was measured by an enzymatic assay and urinary creatinine by a colorimetric method (picric acid), Ca and Mg by atomic absorption spectrophotometry (Model 3110; PerkinElmer, Norwalk, CT, USA), Na and K by flame photometry (IL 943), and phosphorus by colorimetry. Serum-ionized Ca was measured by an ion-selective electrode (ABL705, Radiometer, Copenhagen, Denmark), plasma renin by DiaSorin CLIA method on Liaison automat (DiaSorin, Saluggia, Italy), and aldosterone by liquid chromatography coupled to mass spectrometry with a Waters Xevo TQ-S tandem mass spectrometer (Guyancourt, France).
Animals
Experiments were performed on C57Bl6 wild-type mice. Animals had free access to food (SAFE Laboratory) and tap water. All experiments were conducted in accordance with the institutional guidelines and the recommendations for the care and use of laboratory animals (project authorization number Ce5/2012/084).
Microdissection
Nephron segments from C57Bl6 mice were microdissected from liberase-treated kidneys, as previously reported.16 Briefly, the left kidney was perfused in situ with 6 ml of Hank’s solution supplemented with 1 mM glutamine, 1 mM pyruvate, 0.5 mM MgCl2, 0.1% bovine serum albumin, 20 mM Hepes, and 0.015% liberase (w/v), pH 7.4. Thin pyramids were cut from the kidney and incubated in 0.006% liberase solution for 20–25 min at 30 °C and thoroughly rinsed in microdissection solution supplemented with protease inhibitors cocktail (Complete Tablets, EDTA-free; Roche Diagnostics, Meylan, France).
Messenger RNA extraction and reverse transcription real-time polymerase chain reaction
For nephrons segments, RNAs were extracted from pools of 40–60 segments using the RNeasy micro kit (Qiagen, Courtaboeuf, France) according the manufacturer’s protocol. For mouse tissues, total RNA was extracted from C57Bl6 mice using Tri Reagent (Sigma-Aldrich, Villebon-sur-Yvette, France) according to the protocol described by the manufacturer. RNAs were reverse transcribed using the first-strand complementary DNA (cDNA) synthesis kit for reverse transcription polymerase chain reaction (RT-PCR) (Roche Diagnostics), according to the manufacturer’s protocol. Real-time PCR (qPCR) was performed using a cDNA quantity corresponding to 0.1 mm of the different segments of nephron or 750 ng of mouse tissues with SYBR Green I Master qPCR kit (Roche Diagnostics) on a LightCycler (Roche Diagnostics), according to the manufacturer’s protocol. No DNA was detectable in samples that did not undergo reverse transcription, and in blanks run without cDNA. In each experiment, a standardization curve was made using serial dilutions of a standard cDNA stock solution made from whole kidney. Specific primers were designed using LightCyclerProbeDesign 2 (Roche Diagnostics). Primers used: cldn10 forward 5’- CACAGGCCCTTCAAAACAGT-3’ and reverse 5’-CCCTTACAGAAAGGCACAGC-3’; cldn10b forward 5’-CGATGGCACTGTCATCAC-3’ and reverse 5’-CCTCCGACTTTGGTACATTTCA-3’; and cldn10a forward 5’-GGCTGGCTTTAACATTCGT-3’ and reverse 5’-GAGAGCACCAAACCCTCC-3’.
Cloning of human Claudin-10b
Human CLDN10B was cloned by PCR from a pool of cDNA from human renal biopsy specimens as previously described17 using CLDN10B-specific primers: forward 5’-GCCGCCATGGCTAGCACG-3’and reverse 5’-GACATAAGCATTTTTATCAAACTGTT TTGAAGGG-3’. PCR product was subcloned into pcDNA3.1/V5-HIS TOPO TA (Invitrogen, Villebon-sur-Yvette, France). PCR product insertion was verified by enzymatic digestions with EcoRI and BamHI. Construct was confirmed by sequencing (ATGC).
Site-directed mutagenesis
Methionine at position 1 and serine at position 131 were mutated respectively to threonine and leucine using a QuickChange Multi site-directed mutagenesis kit (Stratagene, Les Ulis, France) on the wild-type CLDN10B construct, and mutations were confirmed by sequencing (GATC Biotech, Mulhouse, France). Primers used to generate c.2T>C CLDN10B mutant: forward 5’-GAATTGCCCTTGCCGCCACGGCTAGCACGG-3’ and reverse 5’-CCGTGCTAGCCGTGGCGGCAAGGGCAATTC-3’ and c.392C>T CLDN10B (c.286C>T CLDN10A) mutant: forward 5’-CTGTCAGGGCTGTGCTTAATGACTGGATGTTCCC-3’ and reverse 5’-GGGAACATCCAGTCATTAAGCACAGCCCTGACAG-3’.
Cell culture and transfections
Mouse kidney thick ascending limb cells were cultured as already described.18, 19 Cells grown for 24 h in 6-well plates were transfected with 4 μg of cDNA construct (CLDN10B, c.2T>C or c.392C>T) using a Magnetofectamine kit (OZ Biosciences, Luminy, France) according to the manufacturer’s protocol. Transiently transfected cells were studied 24 h after transfection for immunocytochemistry or cell surface biotinylation. Empty vector was used as control.
Immunocytochemistry
Twenty-four hours posttransfection, confluent cells were washed with cold PBS2+ (PBS pH8 containing 1 mM MgCl2, and 0.1 mM CaCl2). Cells were then incubated at 4 °C for 40 min in PBS2+ containing 1 mg/ml EZ-Link Sulfo-NHS-LC-biotin (Thermo Fisher Scientific, Villebon-sur-Yvette, France). After cold washing, cells were incubated at 4 °C in 100 mM glycine for 5 min. Then cells were washed with cold PBS2+, fixed with 2% paraformaldehyde in PBS for 20 min at room temperature, permeabilized with 0.1% Triton X-100 for 1 min and incubated with a blocking solution (DAKO, antibody diluent with background-reducing components) for 1 h at room temperature. Cells were incubated with mouse anti-V5 antibody (Abcam, Cambridge, UK) diluted at 1:200 in DAKO for 1 h at room temperature. After washing, cells were incubated in the dark with goat anti-mouse Alexa 488 antibody (Molecular probes, Villebon-sur-Yvette, France) diluted at 1:500 and Cy5-Streptavidine antibody diluted at 1:200 in DAKO. DAPI was used to visualize nucleus. Cells were then mounted with Vectashield. Slides were visualized with a Zeiss LSM 710 laser-scanning microscope using the appropriate filters (Carl Zeiss, Marly-le Roi, France).
Cell surface biotinylation
Confluent cells were placed on ice and washed with cold PBS2+ (PBS pH8 containing 1 mM MgCl2, and 0.1 mM CaCl2). Cells were then incubated at 4 °C for 1 h in PBS2+ containing 1 mg/ml EZ-Link Sulfo-NHS-LC-biotin. Biotinylation was stopped by washing with PBS2+ supplemented with 100 mM glycine. Then they were washed with cold PBS2+ and lysed for 20 to 30 min in 0.2 ml of lysis buffer (100 mM NaCl, 1 mM EDTA, 50 mM Tris, pH 7.4; 1% (vol/vol) Triton X-100; 0.1% SDS) containing protease inhibitors (Complete Tablets, EDTA-free, Roche Diagnostics). Samples were harvested and centrifuged at 10,000g for 10 min at 4 °C. Protein content was measured with protein assay quantification kit (BCA Protein Kit Assay, Pierce, Villebon-sur-Yvette, France). After normalization, aliquots of equal amounts of proteins were taken for total protein fraction, and the rest of proteins were precipitated at 4 °C overnight using Neutravidin-Agarose Resin (Pierce). After overnight incubation, supernatant was removed centrifugation (11,000g for 5 min at 4 °C). Then resins containing biotinylated proteins were washed seven times with cold lysis buffer and centrifuged for 5 min at 11,000g. After last wash, resins were incubated in a solution containing lysis buffer and Laemmli buffer for 5 min at 95 °C. Total protein fractions were also incubated in Laemmli buffer for 5 min at 95 °C. After denaturation, proteins fractions (total and biotinylated) were subjected to sodium dodecyl sulfate polyacrylamide gel electrophoresis and analyzed by western blot using a mouse anti-V5 antibody (Abcam).
Immunofluorescence
Immunofluorescence was performed on paraffin embedded sections of human skin and kidney. Briefly, after deparaffination, antigen retrieval with 10 mM sodium citrate buffer in water bath at 95 °C for 40 min, and blocking (10% goat serum), sections were incubated overnight at 4 °C with monoclonal mouse antibody against Claudin-10, which detects both Claudin-10a and Claudin-10b (Invitrogen; 1:200Ref 415100) and rabbit polyclonal antibody against THP (1:500; Santa Cruz Biotechnology sc-19554, Heidelberg, Germany). Goat anti-mouse Alexa 555 (1:1,000; Invitrogen, Villebon-sur-Yvette, France) and goat anti-rabbit Alexa 488 (1:500; Life Technologies, Villebon-sur-Yvette, France) were used as secondary antibodies.
Na-K-2Cl Cotransporter (NKCC2) staining (rabbit antibody kindly provided by J. Loffing, used at 1:10,000) was conducted using the same protocol on cryosections, and without antigen retrieval. Images were analyzed using a Zeiss AxioVert 200 M microscope (Carl Zeiss).
Results
Clinical case reports
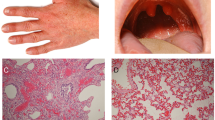
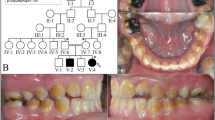
The six reported patients belong to two unrelated families: IV-1, IV-2, IV-3, and IV-4 from family A, and III-4 and V-1 from family B (Figure 1A). All patients were born to consanguineous parents after uneventful pregnancy, labor, and delivery. Patient A-IV-1 is a 24-year-old man. From the age of 9 years, he complained of xerostomia; hypohidrosis of the whole body, including palms and soles; and muscle cramps with minor exercise. At the age of 21, he was referred for heart palpitations, muscle weakness, polydipsia, and polyuria revealing hypokalemia. He presented with xerosis, mild, diffuse ichthyosis, plantar keratoderma, and palmar hyperlinearity (Figure 1B). The nails and hair were normal. Alacrimia was confirmed by Schirmer test strips (Schirmer-Plus blue; Gecis, France) while ocular examination was normal. Xerostomia was confirmed by saliva secretion rate measurement (Figure 1C). Dental examination showed severe enamel wear (Figure 1D). Hearing, growth parameters, and pulmonary function tests were within the normal range. Patients A-IV-2, A-IV-3, and A-IV-4 had workup results similar to those for patient A-IV-1 and showed a considerably similar phenotype.
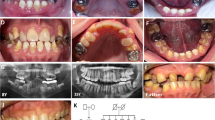
Pedigrees, clinical and histological phenotype of HELIX syndrome. (A) Pedigrees of family A (left panel) and family B (right panel). (B) Skin features of HELIX syndrome. (a) Palmar hyperlinearity (patient A-IV-1); (b) fine white scaly ichthyosis predominately on the axillae (patient B-V-1); (c) plantar keratoderma. (C) Saliva secretion in healthy subjects and three adult patients. Left panel: saliva was collected for 15 minutes under resting conditions in 3 healthy adult subjects (controls) and 3 adult patients (patients A-IV-1, A-IV-2, and B-III-4). The watery and mucous components of the saliva were separated by centrifugation for 10 minutes. In controls, saliva was mainly composed of fluid (red bar), whereas mucus was a minor component (green bar). In patients, the amount of secreted fluid was small in one (A-IV-2) and undetectable in two (A-IV-1 and B-III-4), whereas the amount of mucus was greater than controls. Right panel: the flow of fluid was measured under resting conditions in six healthy adult subjects (controls) and three adult patients (patients A-IV-1, A-IV-2, and B-III-4). The flow of fluid was reduced by 98% in patients compared to controls. (D) Dental findings in HELIX syndrome. Clinical view of patient A-IV-showing poor dental condition with severe enamel wear restored by resin composites on upper incisors, associated with insufficient oral hygiene, generalized gingival inflammation and localized gingival recession on lower central incisors. The panoramic radiograph revealed generalized enamel wear with prosthetic restorations or large carious lesions on premolars and molars. Note that the enamel is still present on the second upper molars and well-detectable on the nonerupted third molar germs (arrows). (E) Histological findings on light microscopy of a skin biopsy specimen. (a) Slight epidermal hyperplasia with a follicular ostial dilatation, full of basket-weave keratin (★) (hematoxylin & eosin, × 400). (b) Abnormally high number of dilated eccrine sweat glands. Epithelial cells of eccrine sweat glands are not cohesive, showing an acantholytic appearance (arrows, hematoxylin & eosin, × 400). Patient A-IV-2. (F) Kidney biopsy specimen. The renal tissue contained both optically normal and fibrotic glomeruli. Fibrosis was extended and predominated at the superficial cortex. Tubules were slightly dilated. No sign of inflammation or vascular damage was visible (patient B-III-4).
Patients B-III-4 and B-V-1 presented with a similar clinical picture. In patient B-III-4 hypokalemia was first discovered at the age of 26, associated with polyuria and polydipsia. Both patients had diffuse and fine white scaly xerosis (Figure 1B).
Histopathologic examination of skin biopsy specimens from patients A-IV-1, A-IV-2, and B-V-1 showed epidermal changes consistent with ichthyosis (Figure 1E). The stratum corneum is slightly thickened. There were a higher number of dilated eccrine sweat glands and the epithelial cells of were noncohesive, with a pseudoacantholytic appearance (Figure 1E).
All patients had a high plasma renin level together with normal to high plasma aldosterone level, normal to low blood pressure and sustained urinary sodium chloride level (Table 1). Plasma potassium was low in adults but normal in children. Plasma bicarbonate and calcium levels were normal. Plasma magnesium level was elevated in five out of six patients. Urinary magnesium excretion was not elevated. Estimated glomerular filtration rate was normal in all patients at first visit, but decreased in patient B-III-4 to 40 ml/min/1.73 m2 with no obvious cause. Renal-CT scan showed normal sized kidneys with no sign of nephrocalcinosis (Supplementary Figure S2A, patient B-III-4). A percutaneous kidney biopsy was performed in patient B-III-4, showing extended fibrosis but no specific damage (Figure 1F). The juxtaglomerular apparatus was more intensely stained by an antirenin antibody (Supplementary Figure S2B) in the patient than in a control.
The genome-wide linkage screen identified two significant runs of homozygosity in the family, on the long arm of chromosome 13 and on the pericentromeric region of chromosome 20. In the region on chromosome 13, CLDN10 singly stood out as the best candidate gene, based on its function in the kidney and its expression pattern. Whole-genome sequencing in patient A-IV-1 and careful examination of the genes in the homozygosity runs identified CLDN10 (chromosome 13q32.1) as the disease-causing gene and excluded all other genes. CLDN10 resequencing in available family A members revealed a missense variation c.386C>T (NM_182848), p.S129L in Claudin-10a (c.392C>T (NM_006984), p.S131L in Claudin-10b) (Figure 2A). Exclusion of any other significant variants, particularly in SLC12A3, CLCNKB, involved in hypokalemia with renal potassium wasting syndromes (Gitelman syndrome MIM 263800; Bartter syndrome MIM 607364), and CLDN16 and CLDN19 genes was provided by whole-genome sequencing for patient A-IV-1. Clinical similarities between the patients prompted us to sequence CLDN10 in patients B-III-4 and B-V-1, which identified a homozygous missense variation, c.2T>C (NM_006984), p.M1T in Claudin-10b, only (Figure 2A). Both mutations changed a highly conserved amino acid (Figure 2A), segregated with the phenotype in available family members, were predicted to be probably damaging by PolyPhen-2 (http://genetics.bwh.harvard.edu/pph2/) and SIFT (http://provean.jcvi.org/genome_submit_2.php), were not present in more than 500 ethnically matched control chromosomes, and were not reported in the 1000 Genomes (http://www.1000genomes.org) and Exome Aggregation Consortium (http://exac.broadinstitute.org/) databases. In family B, the mutation affected the translation initiation codon, which might have been replaced by the next methionine (11 residues downstream) thus translating a truncated protein.
Molecular findings in patients with HELIX syndrome and in wild-type mice. (A) Schematic representation of the human CLDN10 gene and mutations in families A and B. The human CLDN10 is composed of 5 exons; exon 1a and exon 1b are mutually exclusive, generating two alternatively spliced transcripts that encode CLDN10A or CLDN10B. Partial alignments of human (H.s., Uniprot P78369), gorilla (Go.g., Uniprot G3SE82), cow (B.t., Uniprot Q5E9L9), mouse (M.m., Uniprot Q9Z9S6), chicken (Ga.g., Uniprot E1C923) CLDN10B sequences are shown. Mutation in family A is located in exon 2 and changes a highly conserved serine residue (red). Mutation in family B is located in exon 1b and changes a highly conserved methionine residue (red). In human, a methionine residue (green) located downstream could be an alternative site of initiation of translation. (B) Histological findings by immunohistochemistry with CLDN10 antibody on skin biopsy. CLDN10 immunostaining of epithelial cells of eccrine sweat glands appears as a mottled pattern with perinuclear deposition (arrows in a and b, magnification × 400) or as a square labeling of the membrane (arrows in c, magnification × 400) similar to the control (arrows in d, magnification X400) (a, patient A-IV-2; b, patient A-IV-1; c, patient B-III-4; d, control). (C) Cldn10 transcripts levels in mouse tissues and renal tubular segments. Upper panel: quantification of Cldn10 transcripts in the mouse kidney, salivary gland, and skin. Reverse transcription polymerase chain reaction (RT-PCR) analysis of messenger RNAs (mRNAs) encoding Cldn10a or Cldn10b was performed on whole mouse kidney, salivary glands, and skin. Values were corrected for Rlp26 expression. Both Cldn10a and Cldn10b variants were detectable in kidney, salivary gland, and skin. Cldn10b mRNA was more abundant than Cldn10a mRNA in salivary gland and skin. Lower panel: quantification of Cldn10 transcripts along the mouse nephron. RT-PCR analysis of mRNAs encoding Cldn10a or Cldn10b was performed on microdissected renal tubules. Values were corrected for Rlp26 expression. Cldn10a was detected at a significant level in the convoluted (S1) and straight (S3) parts of the proximal tubule. No significant transcript expression was detected in the medullary and cortical thick ascending limbs (mTAL and cTAL, respectively), in the distal convoluted tubule (DCT), the connecting tubule (CNT), the cortical collecting duct (CCD), and outer medullary collecting duct (OMCD). Cldn10b was detected at a significant level in the mTAL and cTAL parts of the renal tubule. No significant transcript expression was detected in S1 and S3, in the distal convoluted tubule (DCT), the connecting tubule (CNT), the cortical collecting duct (CCD), and outer medullary collecting duct (OMCD). (D) Immunolocalization of the Claudin-10 and Na-K-Cl cotransporter2 NKCC2 in the renal cortex. Left panel: double staining with antibodies directed against uromodulin (UMOD, green), specifically expressed in the TAL, and claudin10 (CLDN10, red) in a patient with a nonspecific interstitial chronic kidney disease (top) and in patient B-III-4 bearing a mutation in Claudin-10b (bottom). The upper panel shows that Claudin-10 is normally expressed both in the TAL where it is coexpressed with uromodulin and in another tubular segment not expressing uromodulin (presumably, the proximal tubule); the lower panel shows that Claudin-10 is not detectable in the TAL of patient B-III-4 but still present in the tubular segment not expressing uromodulin. Bar = 50 μm. Right panel: localization of NKCC2 protein in the kidney. The figure shows that the Na-K-Cl cotransporter2 was similarly expressed at the apical domain of the TAL cells in a patient with nonspecific interstitial chronic kidney disease (upper panel) and in patient B-III-4 bearing a CLDN10B mutation. Bar = 50 μm.
Immunohistochemical analysis of skin biopsy specimens showed mottled perinuclear deposition (Figure 2B, patients A-IV-1 and A-IV-2) or normal membrane deposition (Figure 2B, patient B-III-4) of Claudin-10 in the epithelial cells of sweat glands as compared to controls (Figure 2B). In addition to being expressed in the eccrine sweat glands, CLDN10 is also expressed in all layers of the epidermis. The expression of CLDN10 is reduced in the suprabasal layers of the patients as compared to the control (Supplementary Figure S2C). The expression of filaggrin is slightly reduced in the epidermis of all patients (Supplementary Figure S3).
In the mouse, Cldn10a and Cldn10b were expressed in the kidney, salivary glands, and skin (Figure 2C, upper panel). The pattern of expression of Cldn10a and Cldn10b transcripts in the kidney was determined on microdissected mouse tubules; Cldn10a was detected in the proximal tubule (S1 and S3) whereas Cldn10b was detected in the medullary and cortical parts of the thick ascending limb of loop of Henle (TAL); none could be detected beyond the TAL (Figure 2C, lower panel). In the dissected human kidney, CLDN10B expression was also restricted to the TAL.20 We studied Claudin-10 expression in human kidney tissue (patient B-III-4 and a control with nonspecific interstitial nephritis). In the control, Claudin-10 was detected in the TAL, costained by the antiuromodulin antibody, and in another tubular segment, presumably the proximal tubule (Figure 2D and Supplementary Figure S2D). In patient B-III-4 bearing a CLDN10B mutation, the Claudin-10 was no longer detected in the TAL. No difference in cellular localization of the Na-K-2Cl cotransporter NKCC2 was seen between patient B-III-4 and control (Figure 2D).
The functional consequences of the lack of Claudin-10 in the TAL were assessed by studying the response to furosemide in the three adults (A-IV-1, A-IV-2, and B-III-4) and four healthy controls. At baseline, fractional excretion (FE) of chloride was higher and FE of sodium tended to be higher in patients than controls, despite the patients’ secondary hyperaldosteronism. The FE of calcium and magnesium was similar in patients and controls. During furosemide infusion, FE of sodium and chloride increased to similar values in patients and controls. The FE of calcium increased significantly, but was less in patients than controls, whereas FE of magnesium increased more in patients than controls (Figure 3a; Supplementary Table S1). This findings point toward the TAL as the main site of electrolyte transport disturbance and identifies differential consequences for TAL divalent cation handling induced by CLDN10 mutations. In a primary culture of mouse TAL cells, silencing of Cldn10 did not affect phosphorylated Na-K-Cl cotransporter NKCC2 expression but increased the transepithelial resistance (Supplementary Figure S2E). Of note, affected members of family A did not show evidence of disturbed renal proximal transport (hypophosphatemia, metabolic acidosis, or glycosuria).
Functional consequences of CLDN10B mutations. (a) Response to furosemide in three adult patients bearing a CLDN10 mutation and four healthy subjects. Urinary and plasma sodium (upper left panel), chloride (lower left panel), calcium (upper right panel), and magnesium (lower right panel) and creatinine concentrations were measured before (baseline) and during an intravenous infusion of furosemide (5 mg/hour, after an initial intravenous bolus of 5 mg). On baseline, the fractional excretion of chloride was higher (P < 0.05) in patients than in healthy subjects; a nonsignificant trend (P = 0.07) toward a higher fractional excretion of sodium was present in patients. During furosemide infusion, the fractional excretion of sodium and chloride was similar in patients and healthy subjects. Furosemide infusion induced a 14.9 ± 0.5-fold increase in FENa in healthy controls and a 7.2 ± 3.5-fold increase in FENa in the patients (P < 0.01); furosemide infusion induced a 14.0 ± 3.4-fold increase in FECl in healthy controls and a 6.2 ± 2.3-fold increase in FECl in the patients (P < 0.01). On baseline, fractional excretions of calcium and magnesium were similar in patients and healthy subjects. During furosemide infusion, fractional excretion of calcium was lower in patients than in healthy subjects, whereas a nonsignificant trend toward a higher fractional excretion of magnesium was present in patients, compared to healthy subjects. Furosemide infusion induced a 13.7 ± 2.9-fold increase in FECa in healthy controls and a 9.4 ± 2.1-fold increase in FECa in the patients (NS); furosemide infusion induced an 11.0 ± 4.9-fold increase in FEMg in healthy controls and a 6.4 ± 2.9-fold increase in FEMg in the patients (NS). Patients A-IV-1, A-IV-2, B-III-4. Results are medium and interquartile range. (b) Panoramic radiographs in one healthy subject and three patients bearing a CLDN10 mutation. The panoramic radiographs in patients aged 5 to 46 years showed that the enamel formed, as indicated by the observation of tooth germs on dental X-rays, but wore away extremely fast after tooth eruption. (c) Claudin-10b 3D structural modeling. Close-up of the environment of the mutant Claudin-10b (p.S131L) showing the main steric hindrances that the Leu131 side chain generates with neighboring residues Val21 of helix 1 and Ala86 of helix 2 (red disks represent steric clashes). In addition, the -OH of the wild-type Ser131 makes an intrahelical stabilizing h-bond with the carbonyl oxygen of Ser127, bond that is lost in the mutant claudin-10b. (d) Heterologous expression of the c.392T>C CLDN10B variant in mouse kidney thick ascending limb cells. Mouse kidney thick ascending limb cells were transiently transfected by either the wild-type or the mutant c.392C>T CLDN10B complementary DNA (cDNA). The corresponding protein was labeled by an anti-V5 antibody (red, see Methods). Plasma membrane was labeled by biotin and stained by an antibiotin antibody (green). The protein encoded by the wild-type cDNA mainly localized at the plasma membrane, as shown by the colocalization of the protein and biotin. The mutant protein was mainly expressed in the cell, with no detectable expression at the plasma membrane. (e) Suggested model of ion transport across the epithelium of the thick ascending limb of the loop of Henle (TAL), and salivary and sweat glands under normal conditions and in the absence of CLDN10. Under normal conditions (left panel), the TAL (pink) absorbs NaCl via the furosemide-sensitive Na-K-2Cl cotransporter NKCC2 expressed at the apical membrane. Na and Cl exit the cell across the basolateral membrane via the Na,K-ATPase and the chloride channel ClC-Kb, respectively. Active NaCl absorption is an electrogenic process, resulting in a lumen-positive transepithelial potential difference, owing for a passive reabsorption of Na along the paracellular pathway. Salivary and sweat glands epithelia (green) secrete NaCl: NaCl enters the cell across the basolateral membrane via the Na-K-2Cl cotransporter Na-K-Cl cotransporter1 NKCC1; Cl is secreted into the lumen across the apical chloride channel cystic fibrosis transmembrane conductance regulator, thereby creating a lumen-negative transepithelial potential difference; Na exits the cell across the basolateral membrane via the Na,K-ATPase, and is passively secreted into the lumen along the paracellular pathway. In the absence of functional CLDN10 (right panel), the TAL epithelium still actively reabsorbs NaCl but the paracellular diffusion of Na is impaired, resulting in an overall decrease in NaCl absorption; in the salivary and sweat glands, the passive Na secretion is completely abolished.
All patients presented with severe enamel wear that was detectable at all ages. The enamel formed, as indicated by the observation of tooth germs and panoramic radiographs, but wore away quickly after tooth eruption (Figure 3b).
In the 3D model of the structure of Claudin-10b that is based on the 3D crystal structure of mouse claudin-15,14, 15 Leucine-131 clashes sterically with several of the surrounding residues, the most critical being with Valine-21 and Alanine-86 (Figure 3c), thus perturbing the compactness of the helical bundle. Additionally, the intrahelical stabilizing hydrogen bond between the -OH of Serine-131 the carbonyl oxygen of Serine-127 is lost in the mutant CLDN10B (Figure 3c). As a consequence, the newly synthesized Claudin-10b protein is expected to be retained in the cytoplasm of the renal tubular cell without reaching the membrane (Figures 2B and 3d). The p.M1T CLDN10B mutant is similarly retained in the cytoplasm and not expressed at the plasma membrane of renal tubular cells (Figure 2D, Supplementary Figure S2D,F). Similar experiments were done in opossum kidney proximal cells and in rat kidney cells; the mutants were not expressed at the plasma membrane (not shown).
Discussion
We describe a new syndrome featuring hypohidrosis, electrolyte imbalance, lacrimal gland dysfunction, ichthyosis, and xerostomia (HELIX syndrome) caused by biallelic mutations in CLDN10 in six patients belonging to two unrelated families (Figure 1A). The direct effect of the mutation in the first family is a structural destabilization of the 4-helix bundle, translated in the cell as retention of the newly synthesized protein in the cytoplasm given its inability to fold (Figures 2,3 and Supplementary Figure S3). Severity of the disease is related to gland and renal dysfunction.
The xerostomia is associated with polydipsia and high enamel wear thus increasing the risk of caries, tooth sensitivity, dental infections, and early tooth loss. The enamel wear results from the abnormal saliva as previously reported in other syndromes with salivary dysfunction.21 It might be also a consequence of disturbed enamel mineralization (amelogenesis imperfecta) since Cldn10 is expressed in mouse ameloblasts.22
Claudin-10, encoded by CLDN10, is present as two main alternatively spliced variants that differ in the first exon, claudin-10a and claudin-10b.13 Claudin-10a seems to be exclusively expressed in the kidney and uterus while claudin-10b is expressed in most tissues of the mouse with low expression in the liver and higher expression in the kidney.13 Claudin-10, without distinction between isoforms, was detectable in the submandibular salivary glands of the mouse,23 as well as in the cornified layers of the epidermis.24 Utilizing RT-PCR (reverse transcriptase) technique, transcripts of CLDN10 were found in human skin tissue.25 In the kidney, the two isoforms are widely expressed along the nephron, with 10a higher in tubules in the cortex and 10b higher in the medulla.13 At the cellular level, the localization of claudin-10 is restricted to the apical lateral membrane in the jejunum, colon, urinary bladder, and the kidney, but extends basally, all along the lateral membrane in the acinar cells of the exocrine glands, including the pancreas, salivary, and lacrimal glands.26 Claudin-10 is expressed in the epidermis and eccrine sweat glands. Cldn127 and Cldn10 knockout mice die early after birth. Impaired TJ may induce both epidermal thickness and profillagrin processing.28 Clinically, this results in mild ichthyosis and palmar hyperlinearity.8, 29 On the other hand, the abnormal TJ of the epithelial cells of the eccrine sweat glands prevents the secretion of water in the sweat causing anhidrosis with increased skin dryness, intolerance to heat, and inability to regulate body temperature.
Mutations in CLDN10 induce a renal NaCl wasting syndrome at all ages, presumably due to altered NaCl reabsorption in the TAL. Polyuria is inconsistent, and may be due to increased water intake as a result of the xerostomia. Intriguingly, children have no hypokalemia despite secondary hyperaldosteronism, suggesting adaptive mechanisms that may be less effective as they grow older. Hypermagnesemia is present in all patients but more marked in children than adults, and contrasts with normal urinary magnesium excretion, suggesting higher renal tubular reabsorption of magnesium, as reported in the TAL-specific Cldn10 gene deletion mouse model.30 The elevated transepithelial voltage resulting from the absence of functional CLDN10 in the TAL likely explains the trend toward higher Mg reabsorption and hypermagnesemia;30 however, it leaves the lower reabsorption of Ca unexplained. Nephrocalcinosis present in all mice is not reported in the three adult patients.30 Therefore, renal insufficiency in B-III-4 is probably related to another mechanism.
HELIX syndrome illustrates the importance of paracellular transport in the homeostasis of several epithelia.3, 4 The phenotype of patients bearing a biallelic CLDN10 mutation points to several tissues where Na ion transport along the paracellular pathway is critical (Figure 3e). Molecular mechanisms of sweat,31, 32 tears,33 and saliva34 secretion have been partially elucidated. Lacrimal, salivary, and eccrine sweat glands have a similar structure including a secretory part and a duct. In the secretory epithelium, the Na-K-Cl cotransporter NKCC1 and the Na+,K+-ATPase are located at the basolateral plasma membrane. NKCC1 allows electroneutral influx of K+, Cl-, and Na+ into secretory cells. Then, Cl- is secreted across the apical channel cystic fibrosis transmembrane conductance regulator whereas Na+ ions exit the cell via the Na+,K+-ATPase into the interstitium and are secreted through the TJ between secretory cells owed to the transepithelial voltage; water is secreted through the aquaporin 5 channel. Thus the severe phenotype of total absence of fluid in saliva, tears, and sweat might be a consequence of the total loss of Na transport across the TJ. This is consistent with the demonstration that the lack of Cldn10 decreases paracellular Na permeability in mouse TAL30 and increases epithelial resistance in a primary culture of mouse TAL cells (Supplementary Figure S2E).
By contrast, in the TAL, only one half of Na+ reabsorption occurs across the paracellular pathway, the other half being reabsorbed along the transcellular pathway owing to the apical Na-K-Cl cotransporter NKCC2 and the basolateral Na+,K+-ATPase. The transcellular absorption of NaCl is probably preserved in patients bearing CLDN10 mutations, allowing a less severe NaCl wasting than in patients with NKCC2 loss-of-function mutations.35
To date only five claudins are associated with human diseases, namely CLDN1, CLDN14, CLDN16, CLDN19, and CLDN10. Dermatological features are not reported in patients carrying CLDN14, CLDN16, and CLDN19 mutations. Patients carrying biallelic mutations in CLDN1 present with large, scaling, diffuse ichthyosis; scalp hypotrichosis with short, dystrophic, and thick hair; and scarring alopecia at birth.8 The patients in this report have mild scaly ichthyosis predominately on the knees and arms. Involvement of the palms and soles varies from hyperlinearity to focal heel keratoderma. We expect that each of the remaining 22 claudins is associated with a specific phenotype corresponding to its specific pattern of expression in various organs.
References
Weber CR . Dynamic properties of the tight junction barrier. Ann NY Acad Sci 2012;1257:77–84.
González-Mariscal L, Tapia R, Chamorro D . Crosstalk of tight junction components with signaling pathways. Biochim Biophys Acta 2008;1778:729–756.
Krause G, Protze J, Piontek J . Assembly and function of claudins: structure-function relationships based on homology models and crystal structures. Semin Cell Dev Biol 2015;42:3–12.
Brandner JM . Importance of tight junctions in relation to skin barrier function. Curr Probl Dermatol 2016;49:27–37.
Lingaraju A, Long TM, Wang Y, Austin 2nd JR, Turner JR . Conceptual barriers to understanding physical barriers. Semin Cell Dev Biol 2015;42:13–21.
Simon D B, Lu Y, Choate KA et al. Paracellin-1, a renal tight junction protein required for paracellular Mg2+ resorption. Science 1999;285:103–106.
Konrad M, Schaller A, Seelow D et al. Mutations in the tight-junction gene claudin 19 (CLDN19) are associated with renal magnesium wasting, renal failure, and severe ocular involvement. Am J Hum Genet 2006;79:949–57.
Hadj-Rabia S, Baala L, Vabres P et al. Claudin-1 gene mutations in neonatal sclerosing cholangitis associated with ichthyosis: a tight junction disease. Gastroenterology 2004;127:1386–1390.
Wilcox E R, Burton QL, Naz S et al. Mutations in the gene encoding tight junction claudin-14 cause autosomal recessive deafness DFNB29. Cell 2001;104:165–172.
Seelow D, Schuelke M, Hildebrandt F, Nürnberg P . HomozygosityMapper: an interactive approach to homozygosity mapping. Nucleic Acids Res 2009;37:W593–599.
Seelow D, Schuelke M . HomozygosityMapper2012: bridging the gap between homozygosity mapping and deep sequencing. Nucleic Acids Res 2012;40:W516–W520.
Seelow D, Schwarz JM, Schuelke M . GeneDistiller—distilling candidate genes from linkage intervals. PloS One 2008;3:e3874.
Van Itallie CM, Rogan S, Yu A, Vidal LS, Holmes J, Anderson JM . Two splice variants of claudin-10 in the kidney create paracellular pores with different ion selectivities. Am J Physiol Renal Physiol 2006;291:F1288–F1299.
Inai T., Sengoku A, Guan X, Hirose E, Iida H, Shibata Y . Heterogeneity in expression and subcellular localization of tight junction proteins, claudin-10 and -15, examined by RT-PCR and immunofluorescence microscopy. Arch Histol Cyto 2005;68:349–360.
Suzuki H, Nishizawa T, Tani K et al. Crystal structure of a claudin provides insight into the architecture of tight junctions. Science 2014;18;344:304–307.
Cheval L, Pierrat F, Dossat C et al. Atlas of gene expression in the mouse kidney: new features of glomerular parietal cells. Physiol Genomics. 2011;43:161–173.
Disset A, Cheval L, Soutourina O et al. Tissue compartment analysis for biomarker discovery by gene expression profiling. PLoS One 2009;4:e7779.
Azroyan A, Morla L, Crambert G et al. Regulation of pendrin by cAMP: possible involvement in β-adrenergic-dependent NaCl retention. Am J Physiol Renal Physiol 2012;302:F1180.
Bourgeois S, Rossingol P, Grelac F et al. Differentiated thick ascending limb (TAL) cultured cells derived from SV40 transgenic mice express functional apical NHE2 isoform: effect of nitric oxide. Pfulgers Arch 2003;446:672–683.
Chabardès-Garonne D, Mejéan A, Aude JC et al. A panoramic view of gene expression in the human kidney. Proc Natl AcadSci USA 2003;100:13710–113715.
Atar M, Körperich EJ . Systemic disorders and their influence on the development of dental hard tissues: a literature review. J Dent 2010;38:296–306.
Hata M, Kawamoto T, Kawai M, Yamamoto T . Differential expression patterns of the tight junction-associated proteins occludin and claudins in secretory and mature ameloblasts in mouse incisor. Med Mol Morphol 2010;43:102–106.
Hashizume A, Ueno T, Furuse M, Tsukita S, Nakanishi Y, Hieda Y . Expression patterns of claudin family of tight junction membrane proteins in developing mouse submandibular gland. Dev Dyn 2004;;231:425–431.
Troy TC, Rahbar R, Arabzadeh A, Cheung RMK, Turksen K . Delayed epidermal permeability barrier formation and hair follicle aberrations in Inv-Cldn6 mice. Mech Dev 2005;;122:805–819.
Brandner JM, Kief S, Grund C et al. Organization and formation of the tight junction system in human epidermis and cultured keratinocytes. Eur J Cell Biol 2002;;81:253–263.
Inai T, Sengoku A, Guan X, Hirose E, Iida H, Shibata Y . Heterogeneity in expression and subcellular localization of tight junction proteins, claudin-10 and -15, examined by RT-PCR and immunofluorescence microscopy. Arch Histol Cytol 2005;;68:349–360.
Furuse M, Hata M, Furuse K et al. Claudin-based tight junctions are crucial for the mammalian epidermal barrier: a lesson from claudin-1-deficient mice. J Cell Biol 2002;156:1099–1111.
Yuki T, Komiya A, Kusaka A et al. Impaired tight junctions obstruct stratum corneum formation by altering polar lipid and profilaggrin processing. J Dermatol Sci 2013;69:148–158.
Smith FJ, Irvine AD, Terron-Kwiatkowski A et al. Loss-of-function mutations in the gene encoding filaggrin cause ichthyosis vulgaris. Nat Genet 2006;38:337–342.
Breiderhoff T., Himmerkus N, Stuiver M et al. Deletion of claudin-10 (Cldn10) in the thick ascending limb impairs paracellular sodium permeability and leads to hypermagnesemia and nephrocalcinosis. Proc Natl Acad Sci USA 2012;109:14241–14246.
Murota H, Matsui S, Ono E et al. Sweat, the driving force behind normal skin: an emerging perspective on functional biology and regulatory mechanisms. J Dermatol Sci 2015;77:3–10.
Cui CY, Schlessinger D . Eccrine sweat gland development and sweat secretion. Exp Dermatol 2015;24:644–50.
Walcott B, Birzgalis A, Moore LC, Brink PR . Fluid secretion and the Na+ -K+ -2Cl- cotransporter in mouse exorbital lacrimal gland. Am J Physiol Cell Physiol 2005;289:C860–C867.
Baker OJ . Current trends in salivary gland tight junctions. Tissue Barriers 2016;4:e1162348.
Vargas-Poussou R, Feldmann D, Vollmer M et al. Novel molecular variants of the Na-K-2Cl cotransporter gene are responsible for antenatal Bartter syndrome. Am J Hum Genet 1998;62:1332–1340.
Acknowledgements
This study is funded in part by grant NPRP 09-367-3-087 from the Qatar National Research Fund (M.K. and H.E.-S.), as well as the Shafallah Center Foundation, the Fonds National de la Recherche Luxembourg (6903109) (EO), the European Community’s Seventh Framework Programme (305608 EURenOmics), the Swiss National Centre of Competence in Research Kidney Control of Homeostasis (NCCR Kidney.CH) program, the Swiss National Science Foundation (31003A_169850), and the Rare Disease Initiative Zurich (radiz), a clinical research priority program of the University of Zurich, Switzerland. The authors are grateful to Sidra Medical and Research Center for the performance of whole-genome sequencing. We are also grateful to the families who graciously participated in the study.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
The first four authors contributed equally to this work.
The last two authors contributed equally to this work.
Supplementary material is linked to the online version of the paper at
Supplementary information
Rights and permissions
About this article
Cite this article
Hadj-Rabia, S., Brideau, G., Al-Sarraj, Y. et al. Multiplex epithelium dysfunction due to CLDN10 mutation: the HELIX syndrome. Genet Med 20, 190–201 (2018). https://doi.org/10.1038/gim.2017.71
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2017.71
Keywords
This article is cited by
-
The CLDN5 gene at the blood-brain barrier in health and disease
Fluids and Barriers of the CNS (2023)
-
Imputation-powered whole-exome analysis identifies genes associated with kidney function and disease in the UK Biobank
Nature Communications (2023)
-
Integrative genome-wide analyses identify novel loci associated with kidney stones and provide insights into its genetic architecture
Nature Communications (2023)
-
Ichthyosis
Nature Reviews Disease Primers (2023)
-
The role of claudins in homeostasis
Nature Reviews Nephrology (2023)